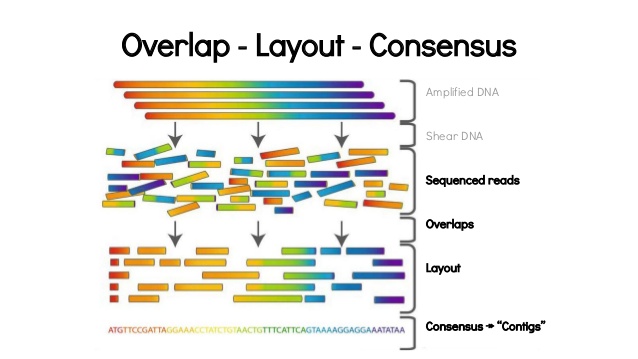
High-throughput pipeline for the de novo viral genome assembly and identification of minority variants from Next-Generation Sequencing of residual diagnostic samples | bioRxiv
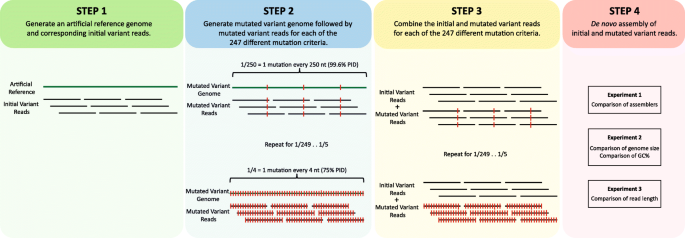
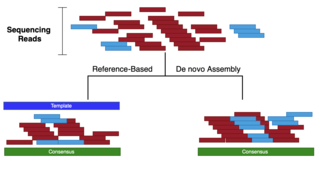
The effect of variant interference on de novo assembly for viral deep sequencing | BMC Genomics | Full Text

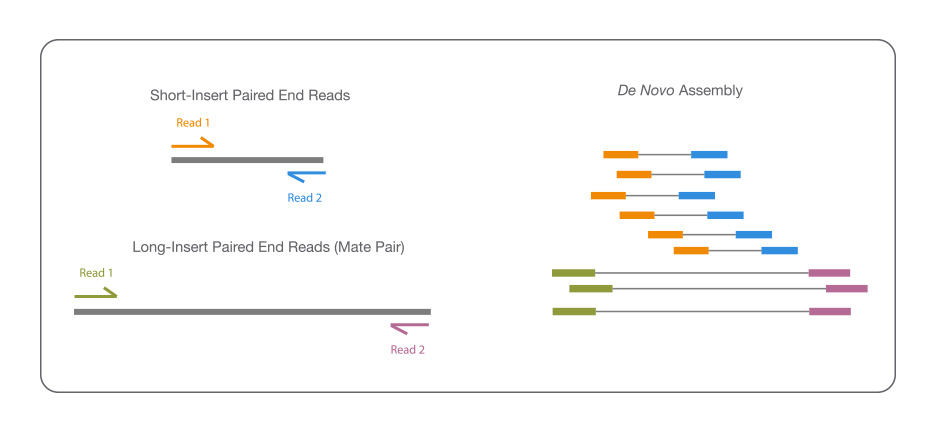
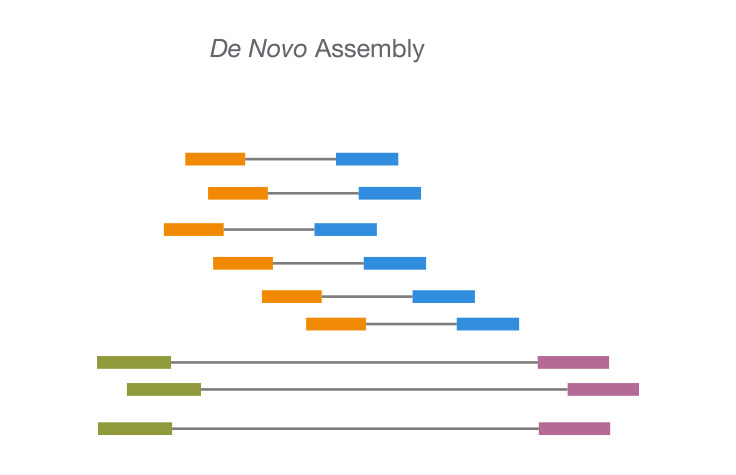
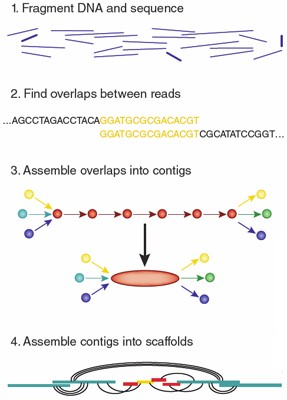
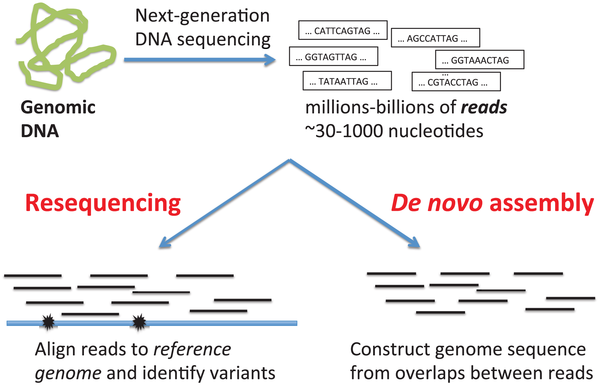
Sequencing from scratch: reference genomes and de novo sequence assembly – HudsonAlpha Institute for Biotechnology

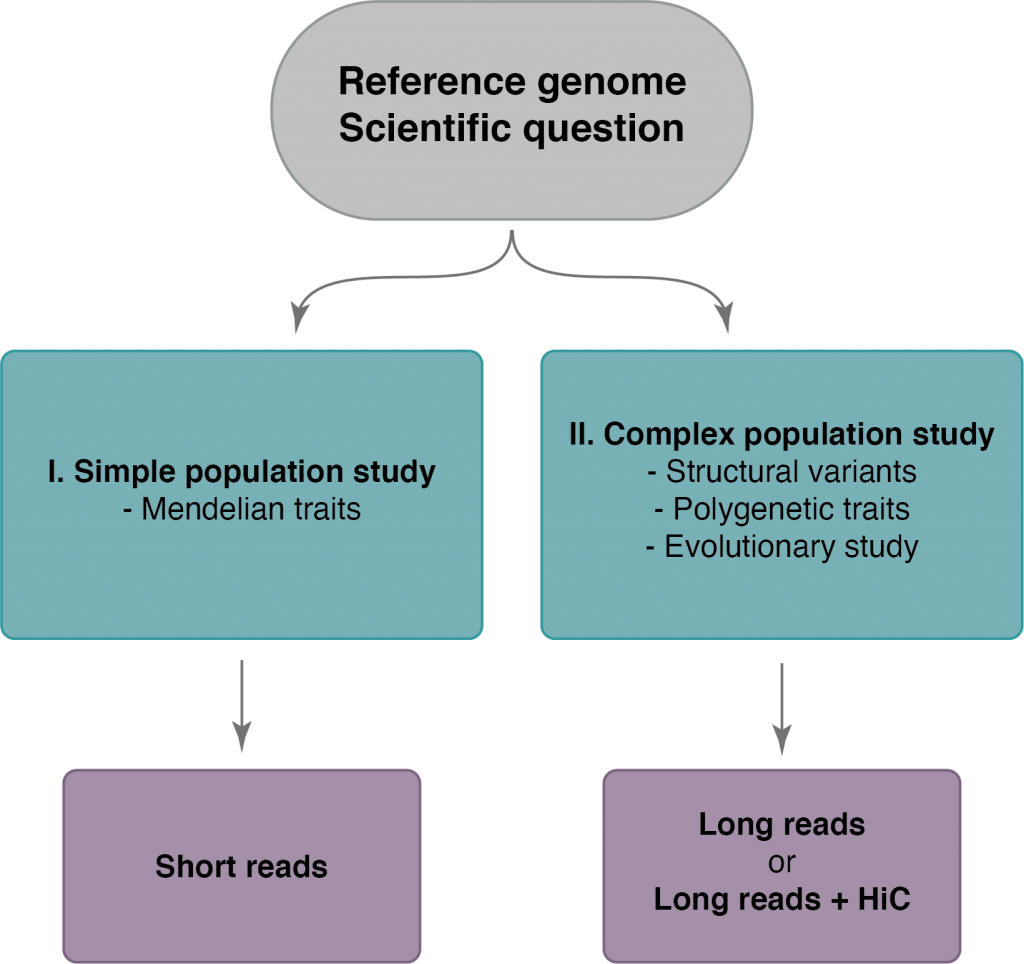
A field guide to whole‐genome sequencing, assembly and annotation - Ekblom - 2014 - Evolutionary Applications - Wiley Online Library
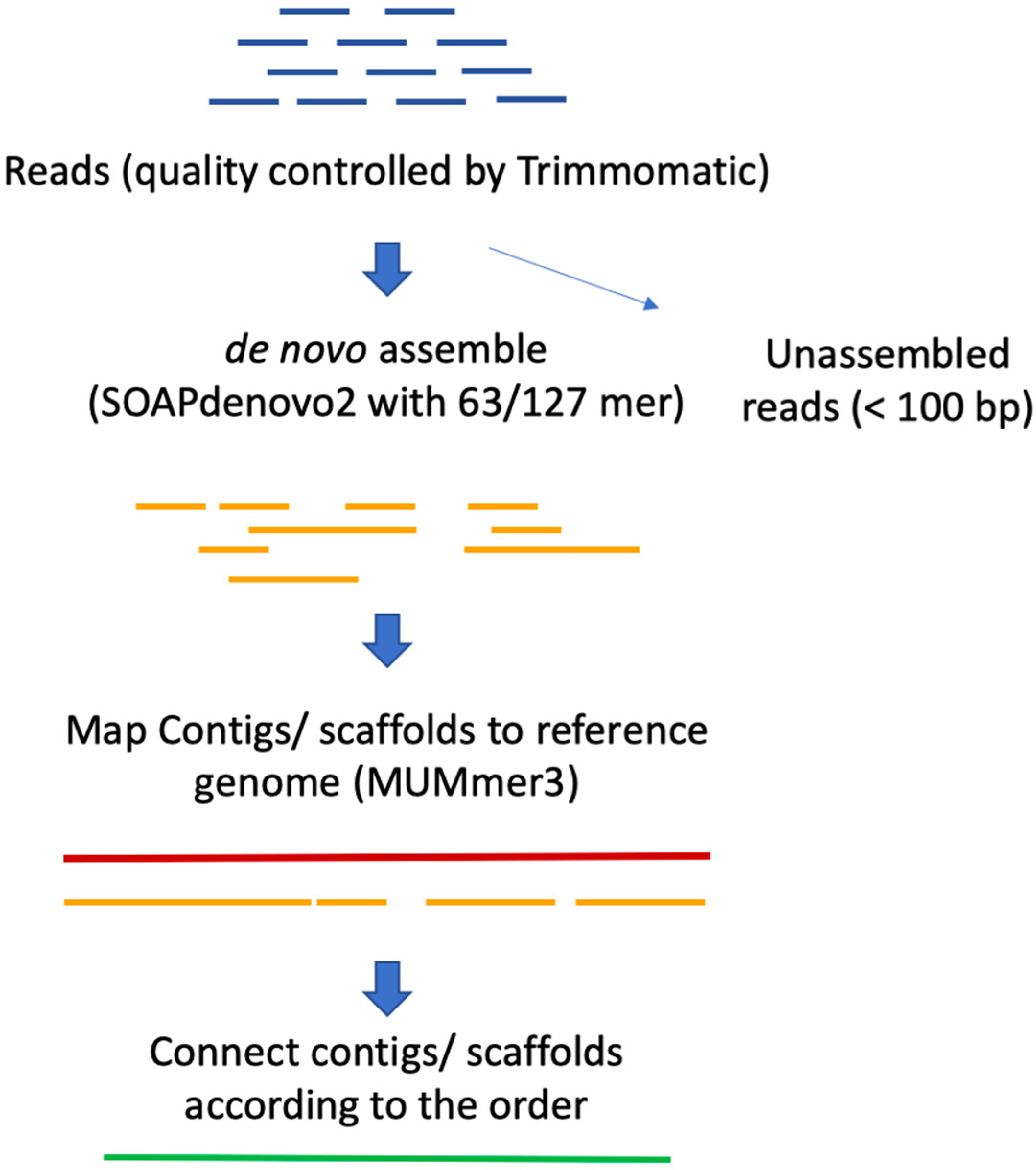
Plants | Free Full-Text | Reference-Guided De Novo Genome Assembly to Dissect a QTL Region for Submergence Tolerance Derived from Ciherang-Sub1
Fine De Novo Sequencing of a Fungal Genome Using only SOLiD Short Read Data: Verification on Aspergillus oryzae RIB40 | PLOS ONE

Sequencing from scratch: reference genomes and de novo sequence assembly – HudsonAlpha Institute for Biotechnology
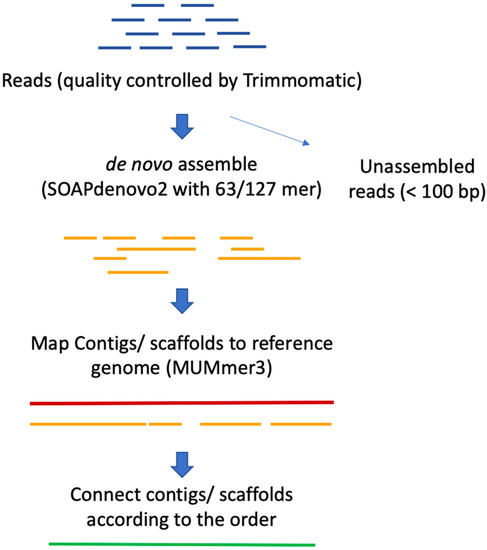
Plants | Free Full-Text | Reference-Guided De Novo Genome Assembly to Dissect a QTL Region for Submergence Tolerance Derived from Ciherang-Sub1




![PDF] RepARK—de novo creation of repeat libraries from whole-genome NGS reads | Semantic Scholar PDF] RepARK—de novo creation of repeat libraries from whole-genome NGS reads | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/b1a85a7249f556c25e875658fa5bd54bb5e1ae08/4-Figure1-1.png)